Theme Lead: Professor Ruth Clifford, Professor Paul Murray
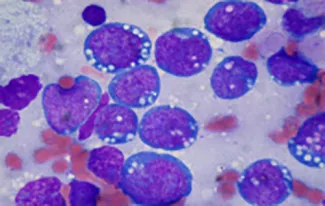
Researchers at the University of Limerick and the University Hospital Limerick are trying to understand the fundamental processes that cause a normal blood cell to become cancerous. If we can understand these processes then we can develop better ways to treat patients and improve cure rates while at the same time reducing the amount of chemotherapy and radiotherapy that the patients receive. We are focused on studying specific types of blood cancer, including different types of lymphoma such as Hodgkin lymphoma and diffuse large B cell lymphoma. (see image below)
A) How does aberrant lysophospholipid signaling contribute to the pathogenesis of Hodgkin's lymphoma and diffuse large B cell lymphoma?
Our studies have so far revealed several S1P-regulated pathogenic pathways and potential vulnerabilities in these pathways that could be exploited therapeutically. In particular, we have identified an oncogenic feed-forward signalling loop that is driven by aberrant S1P signalling and which converges on constitutive activation of the phosphatidylinositol 3 kinase (PI3 K) pathway (Vrzalikova et al., Leukemia 2018 and Vockerodt et al., J Pathol 2019). During the course of these studies we established xenografts of B cell lymphoma. We have used these models to show that S1P is a primary driver of angiogenesis in diffuse large B cell lymphoma and that this effect can be reversed with S1P signalling antagonists, resulting in reduced tumour volumes in vivo (Lupino et al., Leukemia 2019). A further paper is currently in preparation exploring the effects of S1P signalling on macrophage functions in the diffuse large B cell lymphoma microenvironment.
B) How does collagen signalling impact on the pathogenesis of B cell lymphoma?
Collagens are the most abundant non-cellular constituents of tumours. Originally regarded as physical scaffolds, collagens are now known to be ligands for cell surface molecules including the Discoidin domain receptor 1 (DDR1). DDR1 activates cell signalling pathways (e.g. Ras/MAPK) primarily to promote cell adhesion and migratory functions in different cancer types. We have shown that DDR1 can also promote cell survival in lymphoma cells and protect them from chemotherapy. We are now studying if collagen signalling initiated in the tumour microenvironment signals can co-operate with oncogenic signalling pathways to help transform cells.
Application of Machine Learning to Improve Detection of Bone Abnormalities in Multiple Myeloma Patients
Multiple Myeloma (MM) remains an incurable plasma cell malignancy despite improvements in treatment over the last two decades. There are 250 patients newly diagnosed with MM in Ireland annually. Patients undergo a CT Skeletal Survey to assess for the presence of bony lesions associated with MM. Each CT Skeletal Survey generates over a thousand images which are assessed by a Consultant Radiologist to identify bony lesions. Identifying these lesions leads to commencement of treatment to prevent further disease complications and improve overall survival.
There is evidence that identifying patients at high risk of progression to Multiple Myeloma and commencing treatment without delay improves the overall survival of these patients.
Machine learning is a branch of artificial intelligence that provides the ability to a computer programme to automatically learn from experience and apply this learning to detect patterns efficiently. In this project machine learning will be used to teach a computer programme based at the University of Limerick to detect focal bony lesions diagnostic of Multiple Myeloma requiring urgent treatment, and we hypothesise that the diagnostic process will be improved for patients at our centre in terms of faster reporting of results and identification of subtle bony lesions that are hidden within thousands of CT images but detected automatically by the machine learner.
We will characterise novel immune checkpoints in Hodgkin lymphoma: We have already identified the existence of a soluble tumour marker in Hodgkin lymphoma which reflects the presence of tumour-associated macrophages (TAM) in the tumour microenvironment. We have shown that this soluble protein is overexpressed by TAM upon their polarisation in the TME and that the soluble form released by macrophages mediates a novel immune checkpoint.
We are developing new models of lymphoma based on the use of novel retroviruses and lentiviruses to stably transduce primary germinal centre B cells with lymphoma oncogenes. We will use these models to explore how cellular and viral genes co-operate to transform B cells
In the short term we will explore the potential therapeutic benefits in preclinical models, of inhibitors against the enzyme, CAMK1D. We have shown that CAMK1D is overexpressed in some B cell lymphomas. At the same time, we are studying how CAMK1D is regulated and exploring its effects on downstream signalling molecules.
In the medium term we will expand our range of clinical studies to include an exploration of the impact of collagen signalling in a large cohort of patients with Hodgkin lymphoma. We will also begin a clinical study of patients with EBV-associated mucocutaneous (with Dr Graham Taylor and Dr Matthew Pugh) in which we aim to examine if EBV specific immunity both systemically in the blood of patients prior to the development of EBV MCU, as well as in the tissues of patients at the time of diagnosis varies and if it can explain why some patients develop this condition. EBVMCU has recently been recognised by our collaborator, Prof Stefan Dojcinov (with Pugh) as a life threatening complication of Ipilumimab therapy
In the longer term we anticipate that many of the fundamental science studies currently underway will progress to clinical application, work that will be underpinned by our array of pre-clinical models, and our continued access to large, clinically well-characterised, cohorts of patients with lymphomas. In parallel, we will develop the new generation of diagnostic tests that will include companion assays, and enable more precise response prediction and relapse detection.
IMAGES TO BE ADDED
Cader FZ, Vockerodt M, Bose S, Nagy E, Brundler M-A, Kearns P, Murray PG. The EBV oncogene LMP1 protects lymphoma cells from cell death through the collagen mediated activation of DDR1. Blood 2013 122; 4237-4245. SUBJECT OF BLOOD EDITORIAL- Carbone A, Gloghini A. Activated DDR1 increases RS cell survival. Blood 2013; 122: 4152-4.
Vrzalikova K, Ibrahim M, Vockerodt M, Perry T, Margielewska S, Lupino L, Nagy E, Soilleux E, Liebelt D, Hollows R, Last A, Reynolds G, Abdullah M, Curley H, Care M, Krappmann D, Tooze R, Allegood J, Spiegel S, Wei W, Woodman CBJ, Murray PG. S1PR1 drives a feed forward signalling loop to regulate BATF3 and the transcriptional programme of Hodgkin lymphoma cells. Leukemia 2018; 32: 214-223.
Lupino L, Perry T, Margielewska S, Hollows R, Ibrahim I, Care M, Allegood J, Tooze R, Sabbadini R, Reynolds G, Bicknell R, Rudski Z, Hock Y-L, Zanetto U, Wei W, Simmons W, Spiegel S, Woodman CB, Rowe M, Vrzalikova K, Murray PG Sphingosine-1-phosphate signalling drives an angiogenic transcriptional programme in diffuse large B cell lymphoma. Leukemia 2019; May 16. doi: 10.1038/s41375-019-0478-9. [Epub ahead of print]
Vockerodt M, Vrzalikova K, Ibrahim M, Nagy E, Margielewska S, Hollows R, Lupino L, Tooze R, Care M, Simmons W, Schrader A, Perry T, Abdullah M, Foster S, Reynolds G, Dowell A, Rudski Z, Krappmann D, Kube D, Woodman CBJ, Wei W, Taylor GM,. Murray PG. Regulation of S1PR2 by the EBV oncogene LMP1 in aggressive ABC-type diffuse large B cell lymphoma. J Pathol 2019; 248: 142-154.
Alexander L-E, Watters J, Reusch JA, Michelle M, Nepon-Sixt BS, Vrzalikova K, Alexandrow MG, Murray PG, Wright KL. Selective expression of the transcription elongation factor ELL3 in B cells, prior to ELL2, drives proliferation and survival. Mol Immunol 2017;91:8-16
Schrader A, Meyer K, Walther N, Stolz A, Schmidt NM, Hand E, von Bonin F, Ever M, Kohler C, Shirneshan K, Vockerodt M, Klapper W, Szczepanowski M, Murray PG, Bastians H, Trümper L, Spang R, Kube D. Identification of a new gene regulatory circuit by a combined analysis of experimental and clinical high throughput data - B cell receptor mediated suppression of gene expression and G2/M cell cycle aberrations in aggressive lymphoid cancer. Oncotarget 2016; May 7. doi: 10.18632/oncotarget.9219.
Ford CA, Petrova S, Pound JD, Melville L, Paterson M, Ogden CA, Farnworth S, Dumitriu IE, Dunbar DR, Ruckerl D, Allen J, Hume DA, van Rooijen N, Murray PG, Freeman T, Gregory CD. Oncogenic properties of apoptotic tumor cells in aggressive B-cell lymphoma Current Biol 2015; 25: 577-88.
Oldreive CE, Skowronska A, Davies NJ, Parry H, Agathanggelou A, Krysov S, Packham G, Rudzki Z, Cronin L, Vrzalikova K, Murray PG, Odintsova E, Pratt G, Taylor AM, Moss P, Stankovic T. T-cell number and subtype influence the disease course of primary chronic lymphocytic leukaemia xenografts in alymphoid mice. Dis Model Mech. 2015; 8: 1401-12.
Key Publications: Ruth Clifford
- Clifford R, Louis T, Robbe P et al SAMHD1 is mutated recurrently in chronic lymphocytic leukemia and is involved in response to DNA damage Blood. 2014 Feb 13;123(7):1021-31.
-Blakemore SJ, Clifford R, Parker H et al Clinical significance of TP53, BIRC3, ATM and MAPK-ERK genes in chronic lymphocytic leukaemia: data from the randomised UK LRF CLL4 trial. Leukemia. 2020 Feb. doi: 10.1038/s41375-020-0723-2.
-Mark A Catherwood, David Gonzalez, David Donaldson, Ruth Clifford, Ken Mills, Patrick Thornton Relevance of TP53 for CLL diagnostics Journal of Clinical Pathology, Jan 2019 doi.org/10.1136/jclinpath
-Basile Stamatopoulos, Thomas Smith, Emerence Crompot, Karlien Pieters, Ruth Clifford, Marek Mraz et al The light chain IgLV3-21 defines a new poor prognostic subgroup in Chronic Lymphocytic Leukemia: results of a multicenter study Clinical Cancer Research, March 2018, doi: 10.1158/1078-0432.CCR-18-0133
-Burns A, Alsolami R, Becq J, Stamatopoulos B, Timbs A, Bruce D, Robbe P, Vavoulis D, Clifford R, Cabes M, Dreau H, Taylor J, Knight SJL, Mansson R, Bentley D, Beekman R, Martín-Subero JI, Campo E, Houlston RS, Ridout KE, Schuh A. Whole-genome sequencing of chronic lymphocytic leukaemia reveals distinct differences in the mutational landscape between IgHVmut and IgHVunmut subgroups. Leukemia. 2017 Nov 21. doi: 10.1038/leu.2017.311.
-B Stamatopoulos, A Timbs, D Bruce, T Smith, R Clifford , P Robbe , A Burns , D V. Vvoulis, L Lopez, P Antoniou, J Mason, H Dreau, A Schuh Targeted deep sequencing reveals clinically relevant subclonal IgHV rearrangements in chronic lymphocytic leukemia Leukemia. 2017 Apr;31(4):837-845. doi: 10.1038/leu.2016.307. Epub 2016 Oct 31
-Guièze R, Robbe P, Clifford R, de Guibert S, et al Presence of multiple recurrent mutations revealed by targeted NGS confers poor trial outcome of relapsed/refractory CLL Blood. 2015 Oct 29;126(18):2110-7
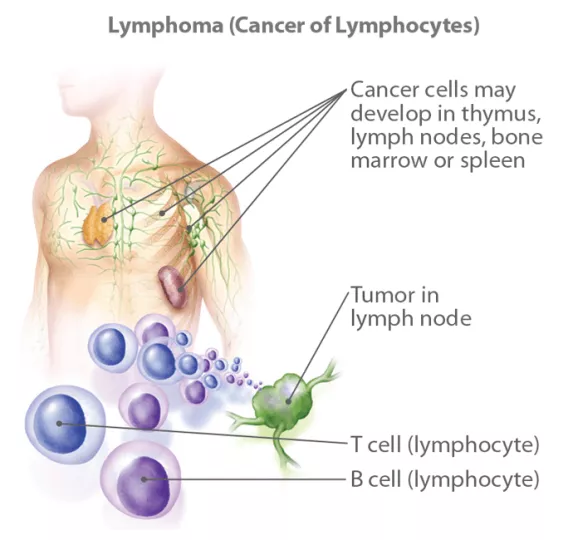
Lymphomas arise from lymphocytes, most commonly from B lymphocytes.